1. Introduction
One of the most prevalent forms of oral cancer is oral squamous cell carcinoma (OSCC), which is recognized as a highly aggressive malignancy with higher morbidity and mortality worldwide. Epidemiological data published by the National Cancer Institute (NCI) states that the overall 5-year survival rate for OSCC is 63% (Fonsêca et al., 2023; Wang et al., 2013). Despite diagnostic strategies being advanced, surgical techniques and treatment outcomes remain poor due to higher rate of recurrence, metastatic progression, and lower response to conventional chemotherapy. In oral cancer, certain dysregulated proteins have been identified as critical in disease progression. Among these, phosphoinositide 3-kinases (PI3Ks) are lipid kinases that phosphorylate phosphatidylinositol-4,5-bisphosphate (PIP2). This results in the generation of PIP3, which recruits and activates downstream effectors, notably AKT, thereby promoting cell survival, proliferation, migration, and angiogenesis. One of the most frequently dysregulated signaling cascades in OSCC is the PI3K/AKT/mTOR pathway (Aggarwal et al., 2019). Owing to the pivotal role in carcinogenesis, PI3K has emerged as a key target for cancer therapeutics. Although PI3K inhibitors have shown strong preclinical efficacy, their clinical utility remains limited, most of which are associated with the central role of the PI3K pathway’s fundamental involvement in normal cellular homeostasis (Vanhaesebroeck et al., 2021). Consequently, combination approaches (PI3K inhibitors + EGFR inhibitors and mTOR inhibitors) are under investigation to overcome compensatory signaling and enhanced therapeutic response (Pons-Tostivint et al., 2017).
In recent years, natural products and traditional herbal formulations have gained considerable interest as complementary and alternative strategies in cancer therapy. Ayurveda, the traditional Indian medical system, has long been using medicinal plants for the management of inflammatory disorders and neoplastic diseases. Berberis aristata DC (Daruharidra) is among the most widely used species attributed to its broad pharmacological properties, including anti-inflammatory, antimicrobial, and anticancer activities (Potdar et al., 2012). Studies demonstrate that the phytochemicals of the plant, particularly alkaloids, flavonoids, glycosides, and saponins, contribute greatly toward its medicinal value. Notably, its bioactive alkaloid, berberine, is well documented for modulating multiple molecular targets involved in tumorigenesis and cancer progression (Wang et al., 2020). A traditional Ayurvedic preparation of Berberis aristata DC, processed in goat’s milk (BaLac), has been prescribed in classical literature for various treatments of the oral cavity, including ulcers and oral cavity tumors (Acharya Vagbhata, 2018). Since the plant has reported therapeutic potential against a spectrum of diseases, including malignancies, such a plant-based formulation may serve as a source of drug-like compounds capable of regulating molecular targets relevant to its traditional indications. Even if the formulation is prescribed in traditional medicine and is known to be effective for oral cancer as indicated, its molecular mechanism in the context of oral carcinogenesis has not been systematically investigated. There are many studies where the integration of traditional knowledge with modern medicine has led to the identification of promising lead molecules of natural origin with favorable therapeutic profiles (Arya et al., 2025; Nadh et al., 2025). In vitro assays, such as MTT-based cytotoxicity testing, apoptosis evaluation, and cell viability analyses, are fundamental tools in anticancer research, as they provide initial quantitative and mechanistic insights into antiproliferative efficacy, cell death pathways, and selective toxicity toward malignant cells (Bathula et al., 2018; Muktapuram et al., 2012).
Concurrently, advances in computational approaches in drug discovery significantly accelerated the identification and prioritization of bioactive molecules. With this rationale, the present study adopted a comprehensive approach that integrates experimental and computational approaches to investigate the anticancer potential of BaLac against OSCC and to identify lead phytoconstituents with promising pharmacological and drug-likeness attributes for the management of OSCC.
Initially, the in vitro cytotoxic and proapoptotic effects of the formulation were assessed in KB cell lines, while its cytocompatibility was tested on normal fibroblast cells. In addition, metabolomic profiling using UHPLC-MS was performed to identify phytochemicals present in BaLac and the parent plant. These identified metabolites were further subjected to molecular docking and comprehensive ADMET screening to evaluate their interaction with the target enzyme PI3K and their potential as safe, orally bioavailable drug candidates.
1.2. Materials and Methods
1.2.1. Plant Collection and Authentication
The plant specimen, along with the root bark, was collected from the Poonch region of Jammu and Kashmir, India. The collected plant materials were identified and taxonomically authenticated by a taxonomist. The leaf and stem specimens were properly labelled, mounted, and preserved on a herbarium sheet, and a corresponding herbarium voucher number was assigned for future reference.
1.2.2. Formulation and plant extract preparation
The BaLac formulation was prepared following its classical Ayurvedic description, with minimal modifications to suit the experimental setup. Initially, the root bark of the plant was thoroughly washed and air-dried for 7 days at 25 ± 2°C. The dried material was coarsely powdered, and distilled water was added in a ratio of 16 parts (v/v) to 1 part (w/w). The mixture was then heated and maintained at a gentle simmer for 2 hours at 90 ± 5°C. The resulting decoction was sequentially filtered through a muslin cloth and Whatman No. 1 filter paper, followed by concentration to one-eighth of its initial volume using a rotary evaporator at 60°C under reduced pressure. The concentrated decoction was blended with an equal volume of pasteurized goat milk (72°C for 15 seconds) and cooked with continuous stirring for approximately 2 hours at 80 ± 5°C to obtain a semi-solid mass. This semisolid material was subsequently dried for 72 hours at 50 ± 2°C in stainless-steel trays, pulverized and passed through a 60-mesh sieve to produce a uniform BaLac powder. The powder was stored in airtight glass containers at 25 ± 2°C and 60 ± 5% relative humidity.
Plant extract preparation: The root bark was thoroughly washed and air-dried for 7 days at 25 ± 2°C. The dried material was coarsely powdered, and distilled water was added 16 parts (v/v) to 1 part (w/w) of the powder. The mixture was then heated and maintained at a gentle simmer for 2 hours at 90 ± 5°C. The resulting decoction was sequentially filtered through a muslin cloth and Whatman No. 1 filter paper, and concentrated to one-eighth of its original volume using a rotary evaporator at 60°C under reduced pressure. The obtained concentrated semisolid extract was further dried for 72 hours at 50 ± 2°C in stainless-steel trays, pulverized and passed through a 60-mesh sieve to yield uniform fine powder. The final powder was stored in airtight glass containers at 25 ± 2°C and 60 ± 5% relative humidity.
1.2.3. In vitro Cytotoxicity, Apoptosis, and Cytocompatibility Analysis
Oral cancer KB cells and fibroblast L929 cells were procured from the National Centre for Cell Science (NCCS), Pune, India. Cells were cultured in Dulbecco’s Modified Eagle’s medium (DMEM) provided with 10% fetal bovine serum (FBS) and 1% antibiotic–antimycotic solution. Cultures were incubated at 37°C in a humidified atmosphere with 5% CO2 and subcultured upon reaching ~70% confluence. Cells at the third passage were used for all experimental assays.
MTT Cytotoxicity Assay (KB Cells): The cytotoxic potential of BaLac against KB oral carcinoma cells was assessed using the Methyl Thiazolyl Tetrazolium (MTT) assay. KB cells were seeded at a density of 5 × 103 cells/well in 96-well microtiter plates and allowed to adhere under standard culture conditions. Following cell adherence, cells were treated with BaLac at various concentrations of 12.5, 25, 50, 100, 250, 500, and 1000 µg/mL and incubated for 24 hours. Subsequently, the treatment medium was replaced with the MTT reagent, and plates were incubated for an additional 4 hours. The resulting formazan crystals were dissolved in DMSO, and the absorbance was recorded at 570 nm using a multimode microplate reader.
AO/EB Apoptosis Staining (KB Cells): For apoptosis assessment, KB cells (2 × 104 cells/well) were seeded into 24-well plates and maintained under standard culture conditions. Following adherence, cells were treated with BaLac at the IC50 concentration for 24 hours, stained with acridine orange–ethidium bromide (AO/EB, 2 µg/mL) dual stain for 15 minutes at 37°C in the dark and washed with 1× PBS. The cells were overlaid with PBS and visualized using a ZOE Fluorescence Cell Imager (Bio-Rad) at 20× magnification under green and red fluorescence channels to distinguish viable, apoptotic, and necrotic populations.
MTT Cytocompatibility Assay (L929 Cells): The cytocompatibility of BaLac was evaluated on L929 fibroblast cells using the MTT assay, following the same experimental procedure employed for KB cells. L929 cells were seeded at 5 × 103 cells/well in 96-well plates, allowed to adhere, and subsequently treated with various concentrations of 12.5, 25, 50, 100, and 200 µg/mL of BaLac for 24 hours. Cell viability was then quantified via MTT assay. All experiments were conducted in triplicate and statistical significance was measured using a paired t-test.
1.2.4. Metabolite Profiling
Phytochemical profiling of both the plant material and the BaLac formulation were performed using ultra-high-performance liquid chromatography coupled with high-resolution mass spectrometry (UHPLC-HRMS). Analyses were conducted on a Dionex UltiMate 3000 RS UHPLC system interfaced with an Orbitrap Eclipse Tribrid Mass Spectrometer (Thermo Fischer Scientific). Chromatographic separation of small molecule constituents was achieved using the Compound Discoverer 3.3.2.31 data processing platform. The mobile phase system consisted of Solvent A (water with 0.1% formic acid), Solvent B (100% acetonitrile with 0.1% formic acid), and Solvent C (100% methanol with 0.1% formic acid). A GOLD C18 reverse-phase HPLC column (inner diameter 2.1 mm, length 100 mm, particle size 1.9 μm) was employed for chromatographic separation of the metabolites. The injection volume was set at 5 µl, and the chromatographic run time was set at 30 minutes, with a flow rate of 0.3 mL/minute. The column effluent was introduced into the mass spectrometer via heated electrospray ionization (H-ESI) interface. Both negative and positive H-ESI modes were used to ionize the compounds, which were then analyzed using an Orbitrap mass analyzer. The MS spectra of the analyzed samples were matched against multiple databases including. Predicted Compositions, mzCloud Search, ChemSpider Search, and MassList Search databases to identify the likely compounds present in the extract. The phytochemical profiles of both the formulation and plant extract were established using a web-based mass spectrometry ecosystem, Global Natural Product Social (GNPS) molecular networking (https://gnps.ucsd.edu/ProteoSAFe/stat6ic/gnps-splash.jsp). Raw data files were converted to GNPS-compatible (“. mzML” format) files using the MSConvert package (Version 3.0.19330, Proteowizard Software Foundation, USA).
1.2.5. Computational Prediction of Phytochemical-Target Interactions in OSCC
Computational predictive analysis was conducted to evaluate the potential regulatory effect of the phytochemicals identified from the medicinal plant and its formulation on the selected molecular target implicated in OSCC.
1.2.6. Preparation of PI3K Protein Structure for Molecular Docking
The X-ray crystallographic structure of human PI3K (PDB ID:1E8Y, resolution: 2.00 Å) was retrieved from the Protein Data Bank (PDB)(https://www.rcsb.org/). A molecular docking study was performed using the Discovery Studio (DS) software suite (Biovia, 2020). Prior to docking, all the crystallographic water molecules were removed manually. “Protein preparation” protocol of DS was used to protonate, to optimize side-chain conformation, and to model loop regions.
1.2.7. Ligand Preparation
The duplicates of identified compounds from the plant and the formulation were removed. Chemical databases, PubChem (https://pubchem.ncbi.nlm.nih.gov/) and ChemSpider (https://www.chemspider.com/), were used to download their three-dimensional structural files. The experimental anticancer drug buparlisib (BKM120) was used as a control. BKM120 is a selective, orally available pan-PI3K inhibitor (Maira et al., 2012).
1.2.8. Molecular docking
The binding pocket of PI3K was defined based on the key amino acid residues associated with inhibitor interactions. The docking grid was centered at the coordinates X = 160.10, Y = 155.44, and Z = 114.21. LibDock protocol of DS was used for virtual screening of the library set of phytochemicals against PI3K. The number of hotspot spheres and docking tolerance was maintained for the default values of 100 and 0.25, respectively. All selected phytochemicals and the reference drug were docked with PI3K.
1.2.9. Drug-likeness and toxicity prediction
Following molecular docking, the pharmacological features of the selected phytochemicals were assessed. Drug-likeness in terms of the oral bioavailability of these molecules was evaluated using the Lipinski filter protocol. Based on Lipinski’s rule, an orally bioavailable drug should have hydrogen bond donors ≤ 5, hydrogen bond acceptors ≤ 10, a molecular weight ≤ 500 Da, and octanol–water partition coefficient ≤ 5 (Lipinski, 2004). Subsequently, the toxicity and pharmacokinetic behavior of the Lipinski-compliant compounds were predicted using the “ADMET” protocol of DS, which is a comprehensive suite that predicts the biological activity of small molecules in terms of pharmacokinetics (Ferreira and Andricopulo, 2019).
1.3. Results
1.3.1. Plant Material Collection and Authentication
The collected plant specimen of Berberis aristata DC was preserved in a herbarium sheet with voucher No: YENDG01, as shown in Figure 1.
1.3.2. In Vitro Cytotoxicity, Apoptosis, and Cytocompatibility Analysis
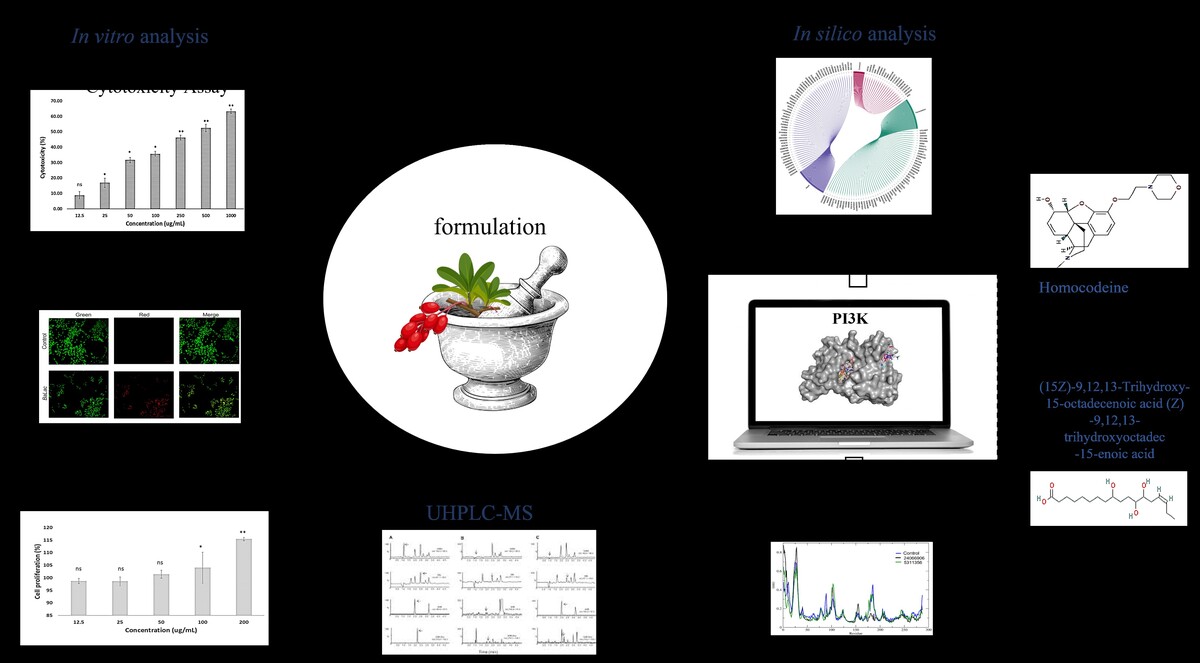
MTT Cytotoxicity Assay: Treatment of KB cells with BaLac resulted in a dose-dependent reduction in cell viability. The IC50 value was determined as 573.01 µg/mL for BaLac. At higher concentrations (250–1000 µg/mL), cell viability decreased significantly compared to that of untreated controls, showing potent antiproliferative effects. The inhibitory activity of BaLac on KB cell proliferation is presented in Figure 2.
Figure 2
Anticancer activity of BaLac in KB cells, as evaluated using MTT assay. X-axis represents concentration of the formulation (µg/mL), ranging from 12.5 to 1000 µg/mL; Y-axis represents cytotoxicity (%); Bars: Represent Mean ± SD of three replicates (n=3). Values are represented as Mean ± SD (n=3). P*<0.05, **p<0.01.
1.3.3. AO/EB Apoptosis Staining (KB Cells)
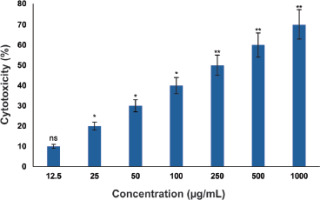
Fluorescence microscopy of AO/EB-stained KB cells revealed distinct apoptotic morphological changes following BaLac treatment at the IC50 concentration. Viable cells exhibited uniform green fluorescence with intact nuclear architecture, whereas early apoptotic cells displayed bright green fluorescence with chromatin condensation. Late apoptotic and necrotic cells displayed orange to red fluorescence, indicating a compromised plasma membrane integrity. Compared to untreated controls, BaLac-treated cultures showed a substantial increase in the apoptotic cell population, confirming the cytotoxicity results obtained in the MTT assay. Representative photomicrographs of control and BaLac-treated KB cells under various channels of the fluorescence imager are shown in Figure 3.
Figure 3
Fluorescence microscopy images showing apoptosis induction in KB cells after treatment with BaLac as evaluated using acridine orange (AO) and ethidium bromide (EB) dual fluorescence staining. Green channel: live cells with intact membranes (AO permeable). Red channel: late apoptotic or necrotic cells with compromised membranes (EB permeable). Merge: Overlay of green and red signals. Control cells predominantly exhibited green fluorescence, indicating high viability. BaLac-treated cells displayed a marked increase in red and yellow/orange fluorescence, indicating significant induction of apoptosis. Magnification: 20×
1.4. Cytocompatibility Analysis
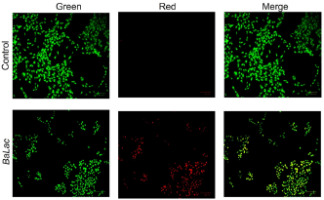
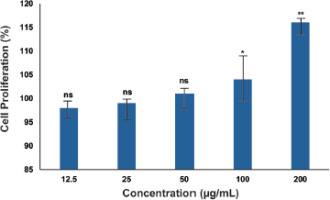
To evaluate the potential cytotoxic effects of BaLac on normal cells, L929 fibroblasts were subjected to an MTT-based viability assay. As illustrated in Figure 4, the viability of L929 fibroblasts treated with BaLac was not compromised. For BaLac, even at the highest tested concentration of 200 µg/mL, cell proliferation was found to be 115.45 ± 0.66%, indicating the cytocompatibility of the formulation on normal cells.
Figure 4
Effect of BaLac on the proliferation of L929 fibroblasts, evaluated using the MTT assay. Cells were treated with different concentrations (12.5–200 µg/mL) of each sample, and proliferation was expressed as a percentage relative to untreated control cells. Data were represented as mean ± SD (n = 3). Statistical significance was determined relative to the control group (*p < 0.05, **p < 0.01). The results indicate that BaLac maintains or enhances fibroblast viability at the tested concentrations, suggesting a noncytotoxic profile toward normal cells.
1.5. Metabolite Profiling
BaLac formulation and the plant extract were analyzed for phytochemical profiling. The analysis identified 3,836 compounds from BaLac, and 4,211 compounds were identified from Berberis aristata DC. The corresponding mass spectrometry datasets have been submitted to MassIVE-Mass Spectrometry Interactive Virtual Environment https (https://massive.ucsd.edu), with accession numbers MSV000096559 (BaLac) and MSV000096558 (Berberis aristata DC), ensuring public accessibility and reproducibility of the analytical results.
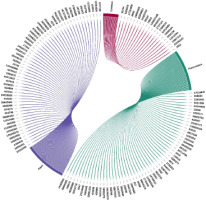
Based on the relative abundance, the top 100 molecules from each sample were selected for docking analysis (supplementary data). After removing duplicates, 169 compounds (86 from the formulation and 83 from the plant) were retained for further analyses. Of these, 17 compounds lacking available structural files were excluded. From the remaining 152 compounds, 20 compounds common to both samples were removed, yielding 132 unique compounds for virtual screening against PI3K. The compounds observed as common and unique to both the formulation and plant are depicted in Figure 5.
Figure 5
Identified compounds from BaLac and Berberis aristata were selected for virtual screening. Green represents compounds from BaLac and, violet represents compounds from Berberis aristata. Common compounds observed in both are represented in magenta. The compounds are represented by their corresponding PubChem/ChemSpider IDs.
1.6. Molecular docking
Molecular docking analysis showed that, out of 132 compounds, 10 compounds demonstrated higher affinity for the selected target, PI3K, than the reference drug. The docking parameters of these compounds are listed in Table 1, and the 2D interaction image is depicted in Figure 6.
Table 1
Docking parameters of top-scored phytochemicals of BaLac and Berberis aristate DC against PI3K.
[i] Note: The top-scoring metabolites identified from BaLac and Berberis aristate DC against PI3K are listed in the table. Ten compounds that showed higher docking scores than the reference drug, and the residues involved in the interactions are shown (Residues forming hydrogen bonds are highlighted).
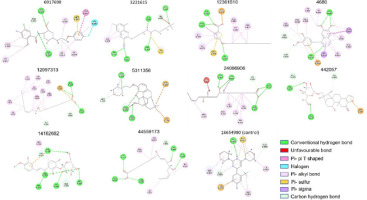
Figure 6
Molecular interactions of identified compounds of BaLac and Berberis aristate with PI3K. The 2D image of intermolecular interactions of top-scored phytochemicals and the control drug are shown. Compounds are represented with their corresponding PubChem IDs. The interacting amino acid residues are labelled and colored according to the type of interactions.
1.7. Drug-likeness and toxicity prediction
All 10 compounds were further screened to assess their pharmacokinetic properties. The parameters of the Lipinski filter and ADMET are listed in Table 2. From the biological activity prediction, it is evident that out of 10 screened compounds, only 2 (homocodeine (PC ID: 5311356), and (15Z)-9,12,13-Trihydroxy-15-octadecenoic acid (PCID: 24066906)) passed all the criteria to act as orally available safe drugs according to the Lipinski filter and ADMET criteria. These compounds were further assessed for their binding stability in terms of thermodynamic parameters using molecular dynamics simulation for 100 ns.
Table 2
Lipinski and ADMET parameters for the selected compounds of BaLac and Berberis aristata.
* PC ID: PubChem ID, HBA: Hydrogen bond acceptor, HBD: Hydrogen bond donor, MW: Molecular weight, ALogP: Octanol-water partition coefficient, Sol: Solubility, Abs: Intestinal absorption, Hep: Hepatotoxicity, CyP2D6: Cytochrome P450 2D6, BBB: Blood–brain barrier, PPB: Plasma protein binding. (Solubility: 0-extremely low, 1-low but possible, 2-possible, 3-good, 4-optimal, 5-too soluble; Intestinal absorption: 0-good, 1-moderate, 2-poor, 3-very poor; BBB level: 0-very high penetrant, 1-high penetrant, 2-medium penetrant, 3-penetrant, 4-undefined (BBB: Blood–brain barrier; CYP2D6: Cytochrome P450 2D6; PPB:Plasma–protein binding).
1.8. Molecular Dynamics simulation
From the simulation data, the compound 5311356 in the formulation showed the highest stability, as indicated by RMSD and RMSF. The RMSD and RMSF plots, the binding modes of the selected compounds, and the control are depicted in Figure 7.
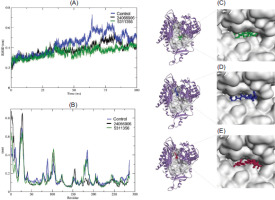
Figure 7
RMSD and RMSF plots and binding mode of selected phytochemicals of BaLac and Berberis aristate. (A) Represents the RMSD plot of 5311356 (green), 24066906 (black), and the control (16654980) (blue). (B) Represents the RMSF plot of 5311356, 24066906, and control (16654980). The binding mode of 5311356, 24066906, and reference drug 16654980 in the binding cleft of PI3K is presented in (C–E), respectively. Docked pose of the compounds in the active cleft of PI3K is shown on the left-hand side. The right-hand side is a zoomed representative of the same in surface view.
2. Discussion
Based on the traditional therapeutic applications and documented pharmacological activities of Berberis aristata DC in diverse disease conditions, including malignancies. The present study explored the anticancer potential of its Ayurvedic formulation, BaLac, against OSCC) through an integrative in vitro and in silico approach. BaLac exhibited a clear concentration-dependent antiproliferative effect on KB cells and significant apoptosis induction, suggesting potential cytotoxicity mediated by programmed cell death rather than nonspecific necrosis.
The antiproliferative effect of the formulation on KB oral cancer cells was assessed using the MTT assay across a concentration range of 12.5 to 1000 µg/mL. The formulation demonstrated a clear dose-dependent anticancer activity against KB cells, from 25 µg/mL onward, and showed maximum effect at 1000 µg/mL with an IC50 value of 573.01 µg/mL. The minimal cytotoxicity (8.84 ± 2.38%) was observed at 12.5 µg/mL, which was not statistically significant compared to the control. Significant cell death (*p < 0.05) was identified from 25 µg/mL (16.88 ± 2.93 %) onward, with further increases at 50 µg/mL (31.69 ± 1.73%) and 100 µg/mL (35.69 ± 1.61 %). Higher concentrations, 250 and 500 µg/mL, produced cytotoxicity of 46.22 ± 1.64% and 52.44 ± 2.50%, respectively, both highly significant (**p < 0.01). The maximum effect (63.30 ± 1.41%) was recorded at 1000 µg/mL(**p < 0.01), indicating potent, concentration- dependent anticancer activity of the formulation against KB cells. The observed IC50 value of 573.01 µg/mL, although relatively high, aligns with values commonly reported for complex herbal or polyherbal matrices in which the concentration reflects the crude formulation rather than isolated active compounds. Such values are within the range considered biologically relevant for initial pharmacological screening of unrefined extracts. Bioactivity-guided fractionation, enrichment of active constituents, or formulation optimization could further enhance potency and reduce the effective concentration required for therapeutic efficacy.
Fluorescence microscopy of AO/EB-stained KB cells provided direct morphological evidence of apoptosis following BaLac treatment. While control cells predominantly exhibited uniform green fluorescence, confirming high viability, BaLac-treated cells exhibited increased red and yellow/orange fluorescence, characteristic of late apoptotic and necrotic populations. These findings are consistent with the MTT assay, further confirming the dose-dependent cytotoxic and apoptosis-inducing effects of the formulation on KB cells.
The effect of BaLac on the proliferation of normal mouse fibroblast (L929) cells was assessed using the MTT assay. From the result, it is clear that BaLac showed no cytotoxicity toward L929 cells at concentrations up to 200 µg/mL. Instead, a slight increase in cell proliferation was observed in a concentration-dependent manner, with the highest proliferation recorded at 200 µg/mL. These findings indicate that the formulation is cytocompatible with normal fibroblasts and may even promote fibroblast growth, in contrast to their potent cytotoxic effects on KB cancer cells.
Taken together, the findings inferred that the reduction in cell viability observed in KB cancer cells is predominantly mediated through apoptosis rather than nonspecific necrosis. Although the maximum inhibition (63.30 ± 1.41%) did not achieve complete cell death, the reduction in cell viability at higher concentrations supports the therapeutic relevance of the formulation. Furthermore, the absence of cytotoxic effects on nonmalignant cells, even at the highest tested concentration, suggests a favorable preliminary safety profile. Nonetheless, future studies should include normal oral epithelial or keratinocyte cell lines to calculate a selectivity index and ensure that BaLac selectively targets malignant cells without affecting normal oral tissues.
The progressive, concentration-dependent inhibitory effect of the formulation may be attributable to the intrinsic bioactive constituents of Berberis aristata DC, as well as to the structurally modified derivatives generated during the formulation process. It is noteworthy that Berberis aristata DC is rich in diverse phytocompounds, particularly berberine, whose protective effect against OSCC has been well established through inhibition of the RAGE/PI3K/AKT/mTOR signaling pathway (Liu et al., 2025). Despite the well-documented anticancer potential, berberine faces major limitations as a promising therapeutic agent due to its poor oral bioavailability, rapid metabolism, and efflux by intestinal P-glycoprotein, which collectively result in very low plasma concentrations after administration. In addition, its short half-life and gastrointestinal toxicity further restrict clinical application(Javed Iqbal et al., 2021; Liu et al., 2010). To address these, many advanced strategies, including liposomal delivery systems and the development of berberine derivatives, are being actively investigated. It is possible that, due to the addition of goat’s milk during the formulation process, structural modifications of the native compounds in Berberis aristata DC might have occurred, leading to the generation of novel derivatives with a promising therapeutic profile. In this context, the present study further evaluated the phytochemical composition of BaLac formulation and compared it with that of Berberis aristata DC. The phytochemical profiling was carried out using the UHPLC-MS method, followed by in silico investigations of the identified constituents against selected key molecular targets of OSCC pathogenesis.
Metabolomic profiling using UHPLC-HRMS revealed a broad and diverse array of phytochemicals in both Berberis aristata and BaLac. The present analysis was qualitative and aimed to identify potential bioactive compounds contributing to the observed effects. Quantitative LC–MS/MS analysis using authentic standards is warranted to establish relative abundance and correlate compound concentration with biological activity. Such quantitative standardization will also aid in defining chemical markers for quality assurance of the formulation.
Dysregulation of the PI3K/AKT/mTOR signaling pathway is strongly implicated in the pathogenesis of various malignancies, including OSCC. This pathway plays a pivotal role in enhancing cancer cell survival, proliferation, and metabolic activity, and has been extensively investigated for its critical involvement in OSCC initiation and progression (Yu et al., 2017). Since the formulation demonstrated antiproliferative activity of the BaLac formulation in cancer cells, the phytochemicals found in Berberis aristata and BaLac were subjected to virtual screening against PI3K to evaluate their potential as PI3K inhibitors.
The in silico assessment provided valuable mechanistic insight, indicating that several BaLac-derived compounds exhibited higher predicted affinity for the PI3K active site than the known inhibitor buparlisib. Molecular docking analysis revealed that, among the 132 screened compounds, 10 compounds exhibited higher binding affinity to PI3K than the known PI3K inhibitor, buparlisib. Among these, five compounds, cisapride (PC ID: 6917698),1-[3-[(3-cyano-6,8-dimethylquinolin-2-yl) amino] propyl]-3-[3-(dimethylamino) propyl], thiourea (PC ID: 3221615), homocodeine (PC ID: 5311356), mascaroside (PC ID: 442057), and okanin 4-methyl ether 3’-glucoside (PC ID: 14162682) were found to be unique for the formulation. The compound, pomalidomide (PC ID: 44571923), was found only in the plant. All the other four identified compounds that demonstrated higher affinity than the control were observed commonly in both plant and formulation (refer to Table 2). Target–ligand interaction profiling (Table 2 and Figure 5) confirmed that these compounds mediate interactions with key amino acids residing in the binding pocket of PI3K, which participate in ATP/ligand binding and catalysis. The observed interacting residues, Lys833, Asp964, Ile879, Tyr867, Val882, and Met953, are the key residues within the ATP-binding pocket. Lys833 (equivalent to Lys802 in canonical numbering) is critical for anchoring ATP’s phosphate groups, and Asp964 (part of the DFG motif) is essential for the catalytic activity (Al Hasan et al., 2023). The repeated engagement of Val882 across multiple top-ranked ligands suggests its role in stabilizing the hydrophobic pocket, even though it is not directly catalytic.
Subsequently, the 10 high-affinity ligands were assessed for oral bioavailability and toxicity to assess their drug-likeness. Among these, two compounds, homocodeine (found only in the formulation) and (15Z)-9,12,13-Trihydroxy-15-octadecenoic acid (found both in the plant and formulation), satisfied all criteria to act as orally available safe drugs. From the binding mode analysis (Figure 7), both compounds occupied the same binding pocket of the reference drug, adopting a similar orientation. Since these compounds exhibited higher affinity and displayed more and stronger engagement with critical residues of the target, they are likely to have superior inhibitory potential compared to the control. Both these compounds were further investigated for their binding stability in terms of thermodynamic parameters using molecular dynamics simulation.
Molecular dynamic simulations performed for 100 ns demonstrated that both homocodeine and (15Z)-9,12,13-Trihydroxy-15-octadecenoic acid remained stably bound within the PI3K active site, correlating with the docking results. RMSD and RMSF analyses collectively suggest that homocodeine exhibited higher structural stability compared to the control and (15Z)-9,12,13-Trihydroxy-15-octadecenoic acid. From the RMSD plot, it is evident that, homocodeine-bound protein complex maintained relatively lower deviation, attaining a study state throughout the 100 ns simulation. For the compound, (15Z)-9,12,13-Trihydroxy-15-octadecenoic acid, a slight deviation of around 0.1 nm was observed around 60–80 ns. In contrast, the reference drug exhibited the highest deviation of around 4 nm, indicating lesser stability. The RMSF plot further supports this observation, where the average residue-wise flexibility for the homocodeine-bound protein complex was consistently lower. In the initial frame, (15Z)-9,12,13-Trihydroxy-15-octadecenoic acid showed the highest initial fluctuation yet converged to a steady state toward the end. The overall results highlight that the selected compounds enhanced the conformational stability of the target protein, minimizing the local flexibility. This suggests the potential of identified compounds as promising ligands.
Collectively, the findings indicate that BaLac exerts selective cytotoxic and proapoptotic effects against OSCC cells, possibly mediated through modulation of PI3K signaling. In the present study, considering the established role of PI3K/AKT/mTOR signaling in OSCC pathogenesis, computational predictive analysis was employed to explore whether phytochemicals identified from BaLac formulation could potentially interact with PI3K. The overall result of in silico analyses revealed that several compounds exhibited favorable predicted binding affinities and stable interactions within the PI3K catalytic pocket compared to the reference inhibitor. However, it is important to emphasize that these observations are derived exclusively from computational prediction and do not provide direct evidence of PI3K enzymatic inhibition or related pathway modulation. Hence, the predicted in silico results require further experimental validation, including in vitro assays and pathway analyses for confirmation. Although further mechanistic and quantitative analyses are required, this study demonstrates a systematic integration of traditional Ayurvedic knowledge with contemporary computational and experimental pharmacology, providing a rational foundation for future translational investigations.
2.1. Study Limitations and Future Perspectives
This study is preliminary in nature, and several limitations should be acknowledged. In the present study, cytotoxicity assessment was confined to a single OSCC cell line (KB), and the observed IC50 value was relatively high, likely reflecting the use of a crude herbal formulation rather than purified active compounds. Evaluation across multiple OSCC cell lines and bioactivity-guided fractionation or isolation of active constituents will be necessary to further establish potency and generalizability. The metabolomic analysis was qualitative, and compound quantification remains to be established. Some matches may represent structurally related natural or synthetic analogues rather than confirmed pharmaceutical entities. This reflects a known limitation of untargeted UHPLC-HRMS–based metabolite annotation, wherein database-driven spectral matching may assign pharmaceutical identities to naturally occurring metabolites or structurally related analogues with identical or near-identical mass and fragmentation features, necessitating further validation with authentic standards. Importantly, although the in silico analyses indicated favorable predicted interactions between selected phytochemicals of the formulation and PI3K, along with promising computationally derived biological activity profiles, no direct biochemical assays, pharmacokinetic evaluations, or experimental toxicity assessments were performed to validate PI3K inhibition or safety. Consequently, definitive mechanistic conclusions cannot be drawn at this stage. Future work should (i) incorporate multiple genetically characterized OSCC models, (ii) include biochemical validation of PI3K signaling inhibition, (iii) compare cytotoxicity with clinically relevant inhibitors such as buparlisib, and (iv) perform MS/MS validation with authentic standards and quantitative LC–MS/MS analysis. In addition, in vivo pharmacokinetic studies and oral toxicity evaluations will be essential to confirm the therapeutic potential and safety of the formulation and its identified compounds.
3. Conclusion
The present study demonstrates that BaLac, an Ayurvedic formulation containing Berberis aristata DC, exhibits dose-dependent cytotoxicity and apoptosis induction in OSCC cells while maintaining cytocompatibility toward normal fibroblasts (Mention the quantitative analysis and justify). Integrative metabolomic and in silico analyses identified key phytoconstituents with favorable PI3K-binding affinity, pharmacokinetic properties, and structural stability. These results support the potential of BaLac as a source of phytochemicals with computationally predicted affinity toward PI3K, providing a rational basis for future experimental validation of PI3K-targeted anticancer mechanisms. Future studies focusing on quantitative metabolite profiling, multicell line screening, kinase inhibition assays, pharmacokinetic and toxicity studies, and in vivo evaluation will be vital to substantiate the therapeutic promise of BaLac in oral cancer management.